Simulating Multiple-Orbital Systems
In this section, we will see how to generate the multiorbital Hamiltonian for a moiré system and compute its band structure. This is essential for understanding the interplay between orbital degrees of freedom and moiré-induced electronic structure. This notebook shows a compact k=2 setup where each hopping can be a 2x2 block.
!!! Here, k denotes the number of orbitals per site and should not be confused with the K-points in momentum space.
1. Setup
We define the lattice once with k=2 and reuse it in all later cells.
import numpy as np
import matplotlib.pyplot as plt
from moirepy import BilayerMoireLattice, TriangularLayer
lattice = BilayerMoireLattice(
latticetype=TriangularLayer,
ll1=2,
ll2=3,
ul1=3,
ul2=2,
n1=1,
n2=1,
k=2,
pbc=True,
)
lattice.generate_connections(inter_layer_radius=1.0)
# Output:
# twist angle = 0.2299 rad (13.1736 deg)
# 19 cells in upper lattice
# 19 cells in lower lattice
2. Build a Block Hamiltonian
Here we mix scalar and matrix-valued inputs:
- In general the t's should be functions that take in certain parameters (as shown in custom_hoppings file) and returns k x k blocks, where k is the number of orbitals per site. But here are some relaxations to the rule.
- If the hopping is supposed to remain constant across the lattice, you can directly input the k x k block without wrapping it in a function.
- If the hopping between any two orbitals is the same, you can directly input a scalar. The code will automatically broadcast it to a full k x k block.
- If the hopping between any two orbitals is the same, but it is different across the lattice, you can input a function that returns a scalar. The code will automatically broadcast it to a full k x k block.
real_ham = lattice.generate_hamiltonian(
tll=0.3,
tuu=0.0,
tul=(
(0.8, 0.5),
(0.5, 0.8),
),
tlu=1.0,
tuself=0.2,
tlself=(
(0.4, 0.1),
(0.1, 0.4),
),
data_type=np.float64, # default is np.complex128
).toarray() # convert to dense array for visualization purposes
print("Hamiltonian shape:", real_ham.shape)
# Output:
# Hamiltonian shape: (76, 76)
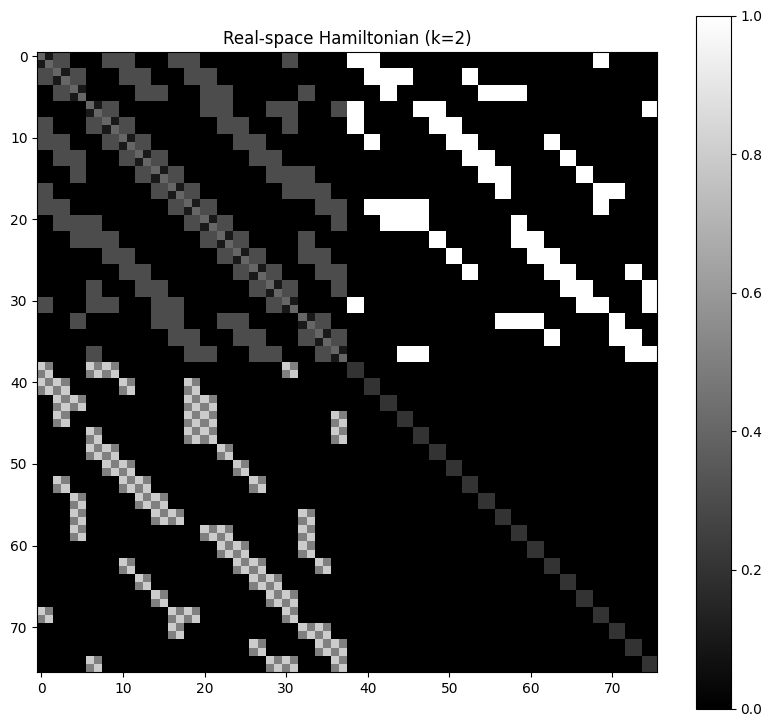
3. Visualize Matrix Structure
plt.figure(figsize=(10, 9))
plt.imshow(real_ham, cmap='gray')
plt.title('Real-space Hamiltonian (k=2)')
plt.colorbar()
# Output:
# <matplotlib.colorbar.Colorbar at 0x7260fbf016a0>
4. Inspect a Small Block
print(real_ham[:16, :16])
# Output:
# [[0.4 0.1 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.3 0.3 0. 0. 0. 0. ]
# [0.1 0.4 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.3 0.3 0. 0. 0. 0. ]
# [0.3 0.3 0.4 0.1 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.3 0.3 0. 0. ]
# [0.3 0.3 0.1 0.4 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.3 0.3 0. 0. ]
# [0. 0. 0.3 0.3 0.4 0.1 0. 0. 0. 0. 0. 0. 0.3 0.3 0.3 0.3]
# [0. 0. 0.3 0.3 0.1 0.4 0. 0. 0. 0. 0. 0. 0.3 0.3 0.3 0.3]
# [0. 0. 0. 0. 0. 0. 0.4 0.1 0.3 0.3 0. 0. 0. 0. 0. 0. ]
# [0. 0. 0. 0. 0. 0. 0.1 0.4 0.3 0.3 0. 0. 0. 0. 0. 0. ]
# [0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.4 0.1 0.3 0.3 0. 0. 0. 0. ]
# [0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.1 0.4 0.3 0.3 0. 0. 0. 0. ]
# [0.3 0.3 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.4 0.1 0.3 0.3 0. 0. ]
# [0.3 0.3 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.1 0.4 0.3 0.3 0. 0. ]
# [0. 0. 0.3 0.3 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.4 0.1 0.3 0.3]
# [0. 0. 0.3 0.3 0.3 0.3 0. 0. 0. 0. 0.3 0.3 0.1 0.4 0.3 0.3]
# [0. 0. 0. 0. 0.3 0.3 0. 0. 0. 0. 0. 0. 0.3 0.3 0.4 0.1]
# [0. 0. 0. 0. 0.3 0.3 0. 0. 0. 0. 0. 0. 0.3 0.3 0.1 0.4]]
Summary
- Setting
k=2promotes each site to a2x2orbital block. - You can combine scalar terms with explicit
(k, k)matrices. - The generated Hamiltonian remains a standard sparse matrix, ready for downstream analysis.